Note
Go to the end to download the full example code.
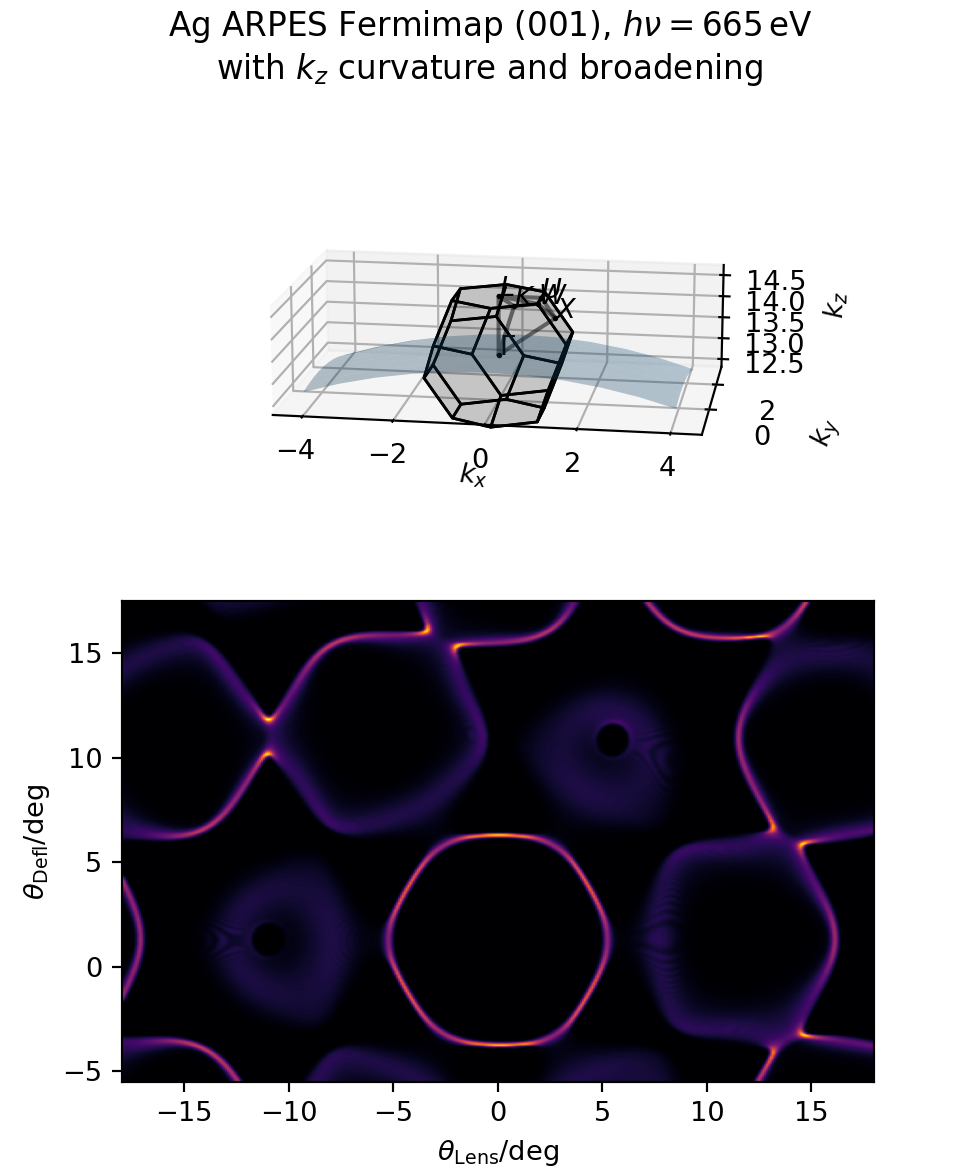
ARPES Fermi surface map simulation¶
/home/docs/checkouts/readthedocs.org/user_builds/fplore/envs/latest/lib/python3.13/site-packages/pymatgen/symmetry/bandstructure.py:179: UserWarning: The input structure does not match the expected standard primitive! The path may be incorrect. Use at your own risk.
kpath = KPathSetyawanCurtarolo(self._structure, symprec, angle_tolerance, atol)
Getting interpolator...
Got interpolator. Doing interpolation...
import numpy as np
from scipy.stats import norm
from fplore import FPLORun
from fplore.util import rot_v1_v2
from fplore.plot import plot_bz
from fplore.arpes import k_arpes, TPS45A1
import matplotlib.pyplot as plt
kz_broadening = True
kz_curvature = True
e_broadening = True
kxky_broadening = True
run = FPLORun("../example_data/Ag")
analyzer_direction = run.high_symm_kpoints['L'] # 111
lens_axis = run.high_symm_kpoints['X'] # 100
# vector rejection of normal to get perpendicular component to surface normal:
lens_axis = lens_axis - (lens_axis @ analyzer_direction) / (analyzer_direction @ analyzer_direction) * analyzer_direction
R = rot_v1_v2(analyzer_direction, [0, 0, 1])
R = rot_v1_v2(R @ lens_axis, [1, 0, 0]) @ R
# usage:
# for multiplying from the left R @ calc coords(3) -> msmt coords(3)
# R.T @ msmt coords(3) -> calc coords(3)
# for multiplying from the right: calc coords(...,3) @ R.T -> msmt coords(...,3)
# msmt coords(...,3) @ R -> calc coords(...,3)
thetacenter, thetawidth = 0, 36
mintheta, maxtheta = thetacenter - thetawidth / 2, thetacenter + thetawidth / 2
theta = np.linspace(np.deg2rad(mintheta), np.deg2rad(maxtheta), 401)
theta2center, theta2width = 6, 23 # 1.3, 12
mintheta2, maxtheta2 = theta2center - theta2width / 2, theta2center + theta2width / 2
theta2 = np.linspace(np.deg2rad(mintheta2), np.deg2rad(maxtheta2), 400)
extent = (mintheta, maxtheta, mintheta2, maxtheta2)
theta, theta2 = np.meshgrid(theta, theta2, indexing='ij')
v0 = 10
workfunc_analyzer = 3.8
e_photon = 665
if kz_broadening:
imfp = 10 # angstroms, see https://doi.org/10.1384/jsa.9.285
Delta_kz = 1 / imfp # standard deviation in reciprocal angstroms (∆o)² = <o²> - <o>²
surface_normal = R @ analyzer_direction # perfectly flat surface perpendicular to analyzer direction
n_kz = 47 # 13
kz_deltas = np.linspace(-3 * Delta_kz, 3 * Delta_kz, n_kz) # shape (N,)
kz_weights_pdf = norm.cdf(kz_deltas, scale=Delta_kz)
kz_limits = np.hstack([[-np.inf], (kz_deltas[1:] + kz_deltas[:-1]) / 2, [np.inf]])
kz_weights = np.array([norm.cdf(u, scale=Delta_kz) - norm.cdf(l, scale=Delta_kz)
for l, u in zip(kz_limits, kz_limits[1:])])
weights = kz_weights.reshape((1, 1, n_kz, 1))
kz_deltas = kz_deltas.reshape((1, 1, n_kz, 1)) * surface_normal.reshape(1, 1, 1, 3) # to shape (1, 1, N, 3)
else:
kz_deltas = 0
weights = 1
k_p_mean = k_arpes(theta=theta, e_photon=e_photon, phi_det=workfunc_analyzer, v0=v0, theta2=theta2, geometry=TPS45A1)
if not kz_curvature:
k_p_mean[:, :, 2] = k_p_mean[:, :, 2].max()
k_p = k_p_mean[:, :, np.newaxis] + kz_deltas # add kz broadening sample points in 3rd dimension
k = k_p @ R # aligned -> calculation (since R == R⁻¹.T)
k_1bz = run.backfold_k(k)
fig = plt.figure(figsize=(5, 6), constrained_layout=True, dpi=196)
#fig.subplots_adjust(wspace=0.6, hspace=0.6)
ax_bz = fig.add_subplot(2, 1, 1, projection='3d')
ax_bz.plot_surface(*(k_p_mean[::40, ::40]).T,
alpha=0.3) # don't draw all the points # - [0, 0, 8] @ run.primitive_lattice.reciprocal_lattice.matrix @ R
midpoint = k.mean(axis=(0, 1, 2))
offset = midpoint @ run.primitive_lattice.reciprocal_lattice.inv_matrix
offset = np.rint(offset)
plot_bz(run, ax_bz, k_points=False, vectors=None, offset=offset, rot=R.T)
ax_bz.view_init(elev=15., azim=-81.)
ax = fig.add_subplot(2, 1, 2)
ax.set_xlabel(r'$\theta_\mathrm{Lens} / \mathrm{deg}$', )
ax.set_ylabel(r'$\theta_\mathrm{Defl} / \mathrm{deg}$')
level_indices = run.band.bands_within(0.2) # (*energy_window)
print('Getting interpolator...')
ip = run.band.get_interpolator(bands=level_indices)
print('Got interpolator. Doing interpolation...')
data = ip(k_1bz)
if not e_broadening:
scale = 0.01 # FWHM, eV
else:
scale = 0.1 # FWHM, eV
scale = scale / 2.355 # FWHM -> std
im = run.band.smooth_overlap(data, e=0, scale=scale, axis=2, weights=weights)
if kxky_broadening:
from scipy.ndimage import gaussian_filter
im = gaussian_filter(im, (1, 1))
ax.imshow(im, extent=extent, origin='lower', aspect='equal', cmap='inferno')
plt.suptitle(f'Ag ARPES Fermimap (001), $h\\nu = {e_photon}\\,\\mathrm{{eV}}$\nwith $k_z$ curvature and broadening')
ax_bz.set_box_aspect([ub - lb for lb, ub in (getattr(ax_bz, f'get_{a}lim')() for a in 'xyz')])
plt.show()
Total running time of the script: (3 minutes 27.997 seconds)